
Group leader: Prof. dr. Pieter Rein ten Wolde
About us
Biochemical networks are the central processing units of the living cell. They allow the cell to sense, transmit, amplify, multiplex, and integrate signals, and even enable it to predict future signals. In our group we use ideas from statistical mechanics and combine analytical theory with innovative computational techniques to elucidate their design principles.
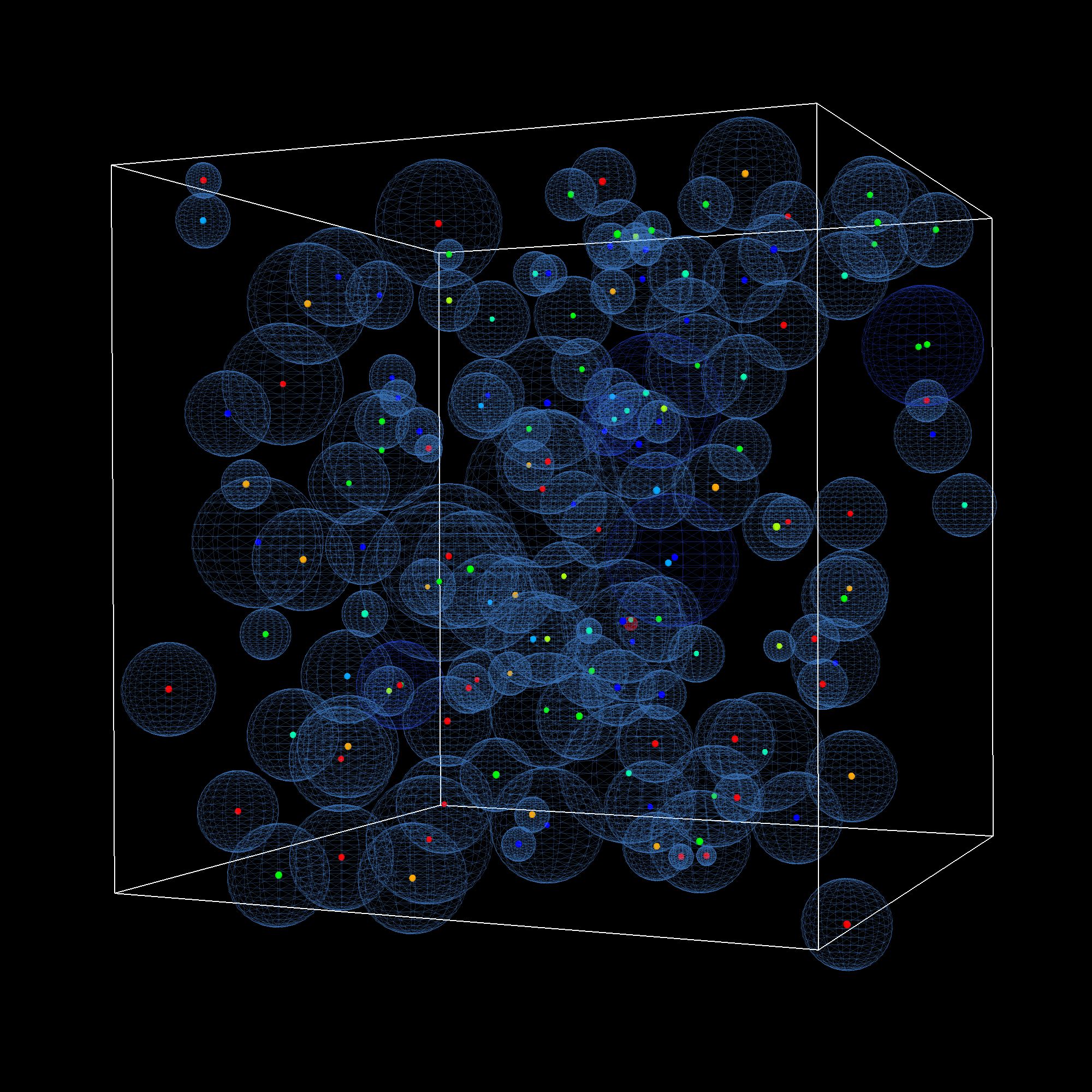

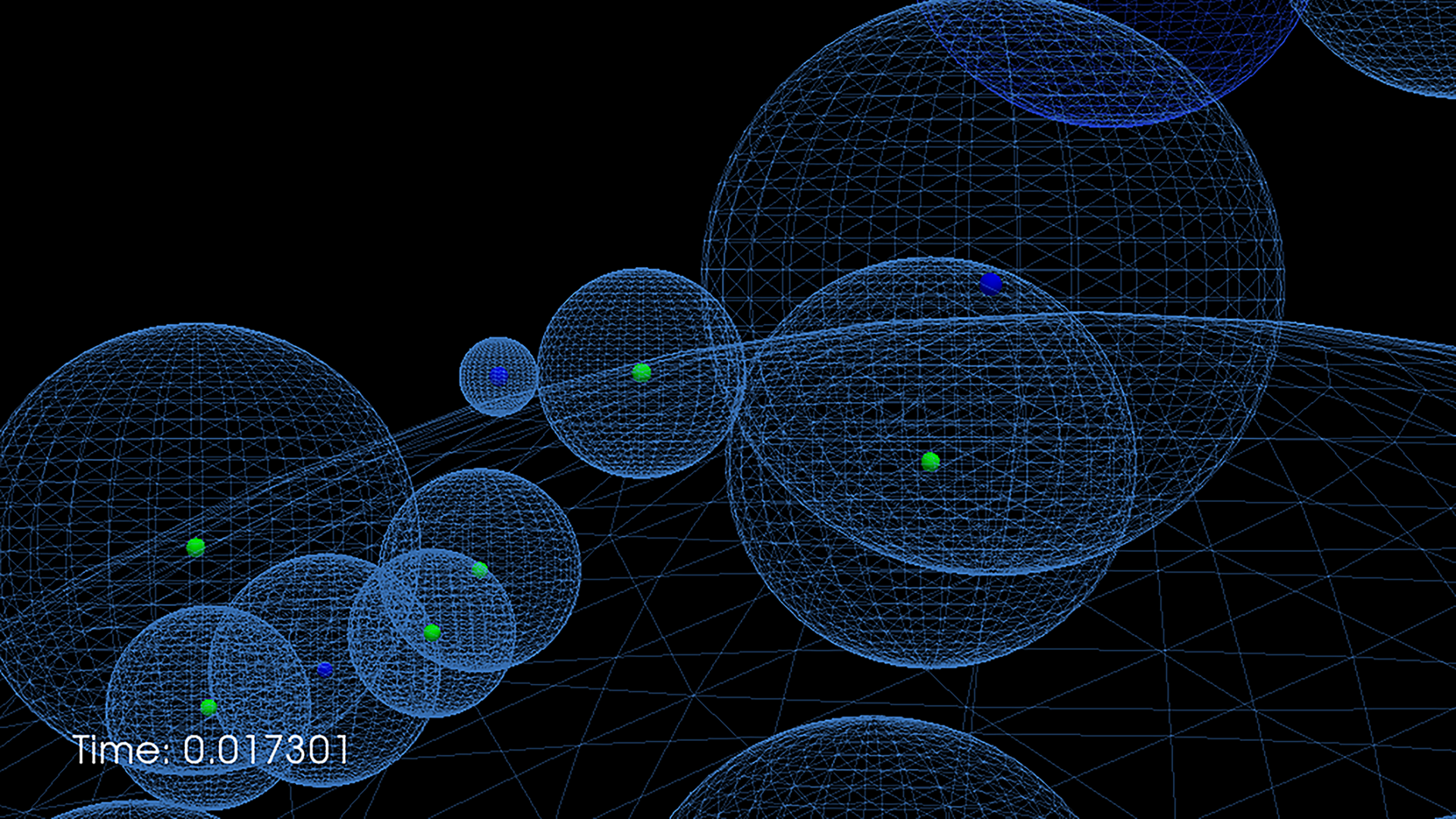
Figure Snapshot of a simulation of a signal transduction pathway (MAPK)
The Biochemical Network group develops and applies new numerical techniques that make it possible to simulate biochemical networks at the particle level in time and space
Recent highlight
New breakthrough in measuring information flow

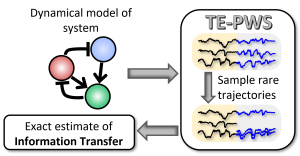
We are proud to share that AMOLF researchers have published a new paper in Physical Review Letters, entitled “Exact Computation of Transfer Entropy with Path Weight Sampling.” The study presents TE-PWS, a computational technique that – for the first time – enables the exact computation of transfer entropy, a central measure in information theory. Transfer entropy is used to quantify how information flows from one part of a system to another, and to detect causal relationships between variables.
Why transfer entropy matters
Group leader Pieter Rein ten Wolde: “This is a breakthrough, because the transfer entropy is the canonical measure for quantifying information transmission and for detecting causal relationships between variables: do the storks deliver babies, or do the storks come in the spring and are babies more often born in the spring?”
From approximation to exact results
Until now, transfer entropy could only be approximated in complex networks, leading to unreliable results. TE-PWS overcomes this by borrowing a concept from statistical physics known as importance sampling, which allows extremely rare fluctuations to be captured efficiently in simulations.
Postdoctoral researcher Avishek Das explains: “Our daily lives rely on complex networks, from the internet and financial markets to ecosystems and the human brain. To understand and control how these networks process information, we need reliable ways to measure information flow. TE-PWS finally makes that possible in any general model.”
The team showed that TE-PWS not only provides exact results but is also as fast – or even faster – than existing approximate methods. This makes it the only reliable and efficient tool available for measuring transfer entropy across a wide range of systems, from cellular reaction networks to stock markets.
Borrowing tools from statistical physics
Beyond its immediate impact in physics and biology, the method could also inspire practical applications. Avishek highlights: “What excites me most is the interdisciplinary nature of this work. We borrowed an idea from physical chemistry to solve an open problem in information theory, and in turn, this solution will help us understand both engineered and biological systems better.”
Potential applications across science and technology
With TE-PWS, scientists and engineers now have a ground-truth method to test their theories, compare approximations, and even design smarter artificial networks. The team envisions applications ranging from understanding brain function to creating bio-inspired materials and autonomous robots.
The code for TE-PWS is openly available through the publication.
Learn more
- If you have questions about this research, please contact Pieter Rein ten Wolde (email: p.t.wolde@amolf.nl).
- This paper was published in Physical Review Letters: “Exact Computation of Transfer Entropy with Path Weight Sampling” by Avishek Das and Pieter Rein ten Wolde.
- Read the full paper: https://doi.org/10.1103/t8z9-ylvg

